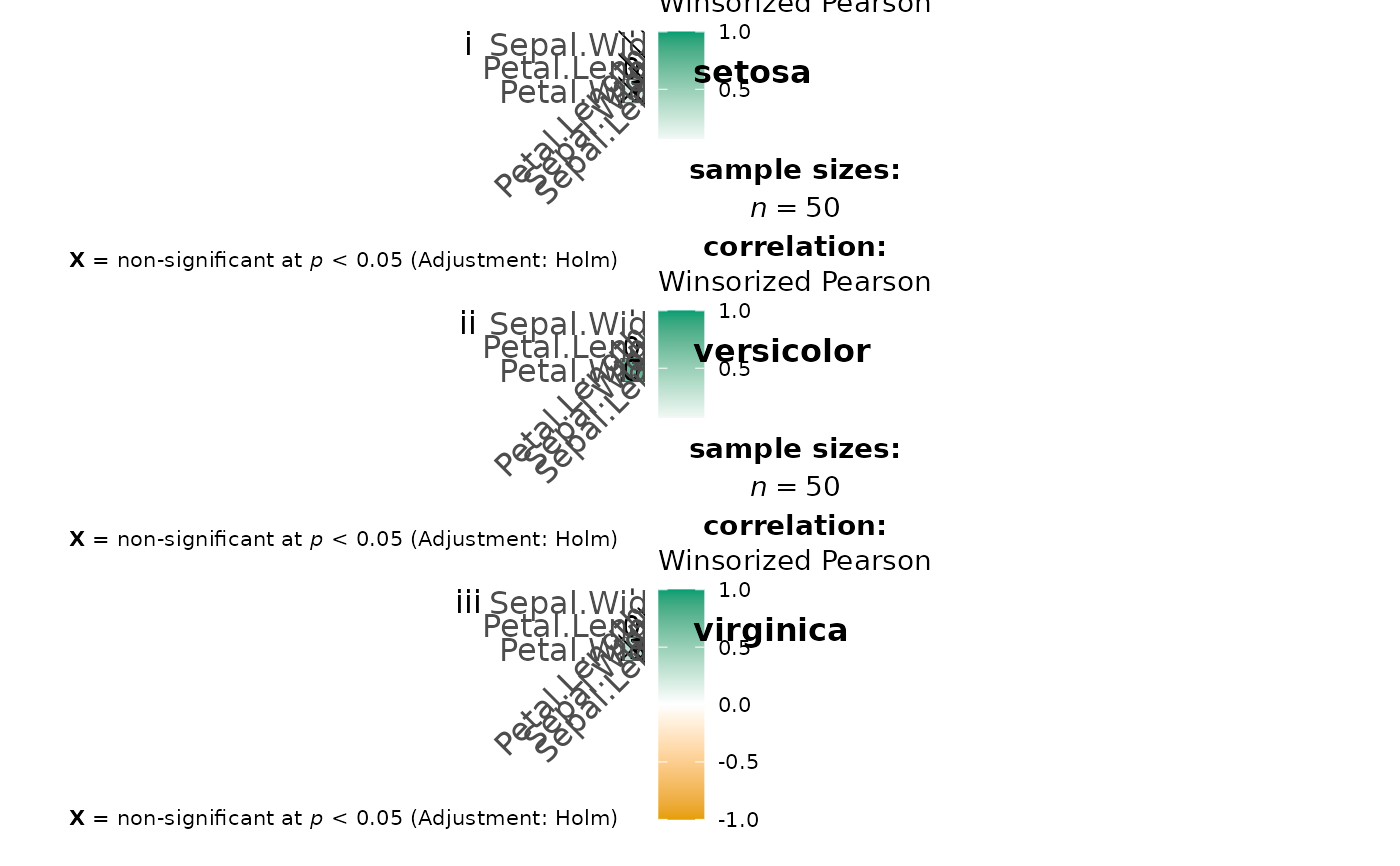
Visualization of a correlalogram (or correlation matrix) for all levels of a grouping variable
Source:R/ggcorrmat.R
grouped_ggcorrmat.RdHelper function for ggstatsplot::ggcorrmat() to apply this function across
multiple levels of a given factor and combining the resulting plots using
ggstatsplot::combine_plots().
Arguments
- data
A data frame from which variables specified are to be taken.
- ...
Arguments passed on to
ggcorrmatcor.varsList of variables for which the correlation matrix is to be computed and visualized. If
NULL(default), all numeric variables fromdatawill be used.cor.vars.namesOptional list of names to be used for
cor.vars. The names should be entered in the same order.partialCan be
TRUEfor partial correlations. For Bayesian partial correlations, "full" instead of pseudo-Bayesian partial correlations (i.e., Bayesian correlation based on frequentist partialization) are returned.matrix.typeCharacter,
"upper"(default),"lower", or"full", display full matrix, lower triangular or upper triangular matrix.sig.levelSignificance level (Default:
0.05). If the p-value in p-value matrix is bigger thansig.level, then the corresponding correlation coefficient is regarded as insignificant and flagged as such in the plot.colorsA vector of 3 colors for low, mid, and high correlation values. If set to
NULL, manual specification of colors will be turned off and 3 colors from the specifiedpalettefrompackagewill be selected.pchDecides the point shape to be used for insignificant correlation coefficients (only valid when
insig = "pch"). Default:pch = "cross".ggcorrplot.argsA list of additional (mostly aesthetic) arguments that will be passed to
ggcorrplot::ggcorrplot()function. The list should avoid any of the following arguments since they are already internally being used:corr,method,p.mat,sig.level,ggtheme,colors,lab,pch,legend.title,digits.typeA character specifying the type of statistical approach:
"parametric""nonparametric""robust""bayes"
You can specify just the initial letter.
digitsNumber of digits for rounding or significant figures. May also be
"signif"to return significant figures or"scientific"to return scientific notation. Control the number of digits by adding the value as suffix, e.g.digits = "scientific4"to have scientific notation with 4 decimal places, ordigits = "signif5"for 5 significant figures (see alsosignif()).conf.levelScalar between
0and1(default:95%confidence/credible intervals,0.95). IfNULL, no confidence intervals will be computed.trTrim level for the mean when carrying out
robusttests. In case of an error, try reducing the value oftr, which is by default set to0.2. Lowering the value might help.bf.priorA number between
0.5and2(default0.707), the prior width to use in calculating Bayes factors and posterior estimates. In addition to numeric arguments, several named values are also recognized:"medium","wide", and"ultrawide", corresponding to r scale values of1/2,sqrt(2)/2, and1, respectively. In case of an ANOVA, this value corresponds to scale for fixed effects.p.adjust.methodAdjustment method for p-values for multiple comparisons. Possible methods are:
"holm"(default),"hochberg","hommel","bonferroni","BH","BY","fdr","none".subtitleThe text for the plot subtitle. Will work only if
results.subtitle = FALSE.captionThe text for the plot caption. This argument is relevant only if
bf.message = FALSE.ggplot.componentA
ggplotcomponent to be added to the plot prepared by{ggstatsplot}. This argument is primarily helpful forgrouped_variants of all primary functions. Default isNULL. The argument should be entered as a{ggplot2}function or a list of{ggplot2}functions.package,paletteName of the package from which the given palette is to be extracted. The available palettes and packages can be checked by running
View(paletteer::palettes_d_names).ggthemeA
{ggplot2}theme. Default value istheme_ggstatsplot(). Any of the{ggplot2}themes (e.g.,ggplot2::theme_bw()), or themes from extension packages are allowed (e.g.,ggthemes::theme_fivethirtyeight(),hrbrthemes::theme_ipsum_ps(), etc.). But note that sometimes these themes will remove some of the details that{ggstatsplot}plots typically contains. For example, if relevant,ggbetweenstats()shows details about multiple comparison test as a label on the secondary Y-axis. Some themes (e.g.ggthemes::theme_fivethirtyeight()) will remove the secondary Y-axis and thus the details as well.
- grouping.var
A single grouping variable.
- plotgrid.args
A
listof additional arguments passed topatchwork::wrap_plots(), except forguidesargument which is already separately specified here.- annotation.args
A
listof additional arguments passed topatchwork::plot_annotation().
Details
For details, see: https://indrajeetpatil.github.io/ggstatsplot/articles/web_only/ggcorrmat.html