Introduction 
{pairwiseComparisons} provides a tidy data friendly way to carry out pairwise comparison tests.
It currently supports post hoc multiple pairwise comparisons tests for both between-subjects and within-subjects one-way analysis of variance designs. For both of these designs, parametric, non-parametric, robust, and Bayesian statistical tests are available.
Installation
| Type | Source | Command |
|---|---|---|
| Release | CRAN | install.packages("pairwiseComparisons") |
| Development | GitHub | remotes::install_github("IndrajeetPatil/pairwiseComparisons") |
Linux users may encounter some installation problems. In particular, the pairwiseComparisons package depends on the PMCMRplus package.
ERROR: dependencies ‘gmp’, ‘Rmpfr’ are not available for package ‘PMCMRplus’
ERROR: dependency ‘pairwiseComparisons’ is not available for package ‘ggstatsplot’This means that your operating system lacks gmp and Rmpfr libraries.
If you use Ubuntu, you can install these dependencies:
sudo apt-get install libgmp3-dev
sudo apt-get install libmpfr-devThe following README file briefly describes the installation procedure: https://CRAN.R-project.org/package=PMCMRplus/readme/README.html
Summary of types of statistical analyses
Following table contains a brief summary of the currently supported pairwise comparison tests-
Between-subjects design
| Type | Equal variance? | Test | p-value adjustment? | Function used |
|---|---|---|---|---|
| Parametric | No | Games-Howell test | ✅ | stats::pairwise.t.test |
| Parametric | Yes | Student’s t-test | ✅ | PMCMRplus::gamesHowellTest |
| Non-parametric | No | Dunn test | ✅ | PMCMRplus::kwAllPairsDunnTest |
| Robust | No | Yuen’s trimmed means test | ✅ | WRS2::lincon |
| Bayesian | NA |
Student’s t-test | NA |
BayesFactor::ttestBF |
Within-subjects design
| Type | Test | p-value adjustment? | Function used |
|---|---|---|---|
| Parametric | Student’s t-test | ✅ | stats::pairwise.t.test |
| Non-parametric | Durbin-Conover test | ✅ | PMCMRplus::durbinAllPairsTest |
| Robust | Yuen’s trimmed means test | ✅ | WRS2::rmmcp |
| Bayesian | Student’s t-test | NA |
BayesFactor::ttestBF |
Examples
Here we will see specific examples of how to use this function for different types of
- designs (between or within subjects)
- statistics (parametric, non-parametric, robust, Bayesian)
- p-value adjustment methods
Between-subjects design
# for reproducibility
set.seed(123)
library(pairwiseComparisons)
library(statsExpressions) # for data
# parametric
# if `var.equal = TRUE`, then Student's *t*-test will be run
pairwise_comparisons(
data = ggplot2::msleep,
x = vore,
y = brainwt,
type = "parametric",
var.equal = TRUE,
paired = FALSE,
p.adjust.method = "bonferroni"
)
#> # A tibble: 6 x 6
#> group1 group2 p.value test.details p.value.adjustment
#> <chr> <chr> <dbl> <chr> <chr>
#> 1 carni herbi 1 Student's t-test Bonferroni
#> 2 carni insecti 1 Student's t-test Bonferroni
#> 3 carni omni 1 Student's t-test Bonferroni
#> 4 herbi insecti 1 Student's t-test Bonferroni
#> 5 herbi omni 0.979 Student's t-test Bonferroni
#> 6 insecti omni 1 Student's t-test Bonferroni
#> label
#> <chr>
#> 1 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 2 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 3 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 4 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 5 list(~italic(p)[Bonferroni-corrected]==0.979)
#> 6 list(~italic(p)[Bonferroni-corrected]==1.000)
# if `var.equal = FALSE`, then Games-Howell test will be run
pairwise_comparisons(
data = ggplot2::msleep,
x = vore,
y = brainwt,
type = "parametric",
var.equal = FALSE,
paired = FALSE,
p.adjust.method = "bonferroni"
)
#> # A tibble: 6 x 11
#> group1 group2 statistic p.value alternative method distribution
#> <chr> <chr> <dbl> <dbl> <chr> <chr> <chr>
#> 1 carni herbi 2.17 1 two.sided Games-Howell test q
#> 2 carni insecti -2.17 1 two.sided Games-Howell test q
#> 3 carni omni 1.10 1 two.sided Games-Howell test q
#> 4 herbi insecti -2.41 1 two.sided Games-Howell test q
#> 5 herbi omni -1.87 1 two.sided Games-Howell test q
#> 6 insecti omni 2.19 1 two.sided Games-Howell test q
#> p.adjustment test.details p.value.adjustment
#> <chr> <chr> <chr>
#> 1 none Games-Howell test Bonferroni
#> 2 none Games-Howell test Bonferroni
#> 3 none Games-Howell test Bonferroni
#> 4 none Games-Howell test Bonferroni
#> 5 none Games-Howell test Bonferroni
#> 6 none Games-Howell test Bonferroni
#> label
#> <chr>
#> 1 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 2 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 3 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 4 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 5 list(~italic(p)[Bonferroni-corrected]==1.000)
#> 6 list(~italic(p)[Bonferroni-corrected]==1.000)
# non-parametric
pairwise_comparisons(
data = ggplot2::msleep,
x = vore,
y = brainwt,
type = "nonparametric",
paired = FALSE,
p.adjust.method = "none"
)
#> # A tibble: 6 x 11
#> group1 group2 statistic p.value alternative method
#> <chr> <chr> <dbl> <dbl> <chr> <chr>
#> 1 carni herbi 0.582 0.561 two.sided Dunn's all-pairs test
#> 2 carni insecti 1.88 0.0595 two.sided Dunn's all-pairs test
#> 3 carni omni 1.14 0.254 two.sided Dunn's all-pairs test
#> 4 herbi insecti 1.63 0.102 two.sided Dunn's all-pairs test
#> 5 herbi omni 0.717 0.474 two.sided Dunn's all-pairs test
#> 6 insecti omni 1.14 0.254 two.sided Dunn's all-pairs test
#> distribution p.adjustment test.details p.value.adjustment
#> <chr> <chr> <chr> <chr>
#> 1 z none Dunn test None
#> 2 z none Dunn test None
#> 3 z none Dunn test None
#> 4 z none Dunn test None
#> 5 z none Dunn test None
#> 6 z none Dunn test None
#> label
#> <chr>
#> 1 list(~italic(p)[uncorrected]==0.561)
#> 2 list(~italic(p)[uncorrected]==0.060)
#> 3 list(~italic(p)[uncorrected]==0.254)
#> 4 list(~italic(p)[uncorrected]==0.102)
#> 5 list(~italic(p)[uncorrected]==0.474)
#> 6 list(~italic(p)[uncorrected]==0.254)
# robust
pairwise_comparisons(
data = ggplot2::msleep,
x = vore,
y = brainwt,
type = "robust",
paired = FALSE,
p.adjust.method = "fdr"
)
#> # A tibble: 6 x 10
#> group1 group2 estimate conf.level conf.low conf.high p.value
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 carni herbi -0.0323 0.95 -0.248 0.184 0.790
#> 2 carni insecti 0.0451 0.95 -0.0484 0.139 0.552
#> 3 carni omni 0.00520 0.95 -0.114 0.124 0.898
#> 4 herbi insecti 0.0774 0.95 -0.133 0.288 0.552
#> 5 herbi omni 0.0375 0.95 -0.182 0.257 0.790
#> 6 insecti omni -0.0399 0.95 -0.142 0.0625 0.552
#> test.details p.value.adjustment
#> <chr> <chr>
#> 1 Yuen's trimmed means test FDR
#> 2 Yuen's trimmed means test FDR
#> 3 Yuen's trimmed means test FDR
#> 4 Yuen's trimmed means test FDR
#> 5 Yuen's trimmed means test FDR
#> 6 Yuen's trimmed means test FDR
#> label
#> <chr>
#> 1 list(~italic(p)[FDR-corrected]==0.790)
#> 2 list(~italic(p)[FDR-corrected]==0.552)
#> 3 list(~italic(p)[FDR-corrected]==0.898)
#> 4 list(~italic(p)[FDR-corrected]==0.552)
#> 5 list(~italic(p)[FDR-corrected]==0.790)
#> 6 list(~italic(p)[FDR-corrected]==0.552)
# Bayesian
pairwise_comparisons(
data = ggplot2::msleep,
x = vore,
y = brainwt,
type = "bayes",
paired = FALSE
)
#> # A tibble: 6 x 18
#> group1 group2 term estimate conf.level conf.low conf.high pd
#> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 carni herbi Difference 0.376 0.95 -0.525 1.33 0.800
#> 2 carni insecti Difference -0.0348 0.95 -0.127 0.0425 0.818
#> 3 carni omni Difference 0.0440 0.95 -0.139 0.239 0.693
#> 4 herbi insecti Difference -0.394 0.95 -1.61 0.775 0.758
#> 5 herbi omni Difference -0.362 0.95 -1.06 0.345 0.859
#> 6 insecti omni Difference 0.0762 0.95 -0.153 0.339 0.732
#> rope.percentage prior.distribution prior.location prior.scale bf10
#> <dbl> <chr> <dbl> <dbl> <dbl>
#> 1 0.171 cauchy 0 0.707 0.540
#> 2 0.134 cauchy 0 0.707 0.718
#> 3 0.236 cauchy 0 0.707 0.427
#> 4 0.166 cauchy 0 0.707 0.540
#> 5 0.162 cauchy 0 0.707 0.571
#> 6 0.161 cauchy 0 0.707 0.545
#> method log_e_bf10 expression label
#> <chr> <dbl> <list> <chr>
#> 1 Bayesian t-test -0.617 <language> list(~log[e](BF['01'])==0.62)
#> 2 Bayesian t-test -0.332 <language> list(~log[e](BF['01'])==0.33)
#> 3 Bayesian t-test -0.851 <language> list(~log[e](BF['01'])==0.85)
#> 4 Bayesian t-test -0.616 <language> list(~log[e](BF['01'])==0.62)
#> 5 Bayesian t-test -0.560 <language> list(~log[e](BF['01'])==0.56)
#> 6 Bayesian t-test -0.606 <language> list(~log[e](BF['01'])==0.61)
#> test.details
#> <chr>
#> 1 Student's t-test
#> 2 Student's t-test
#> 3 Student's t-test
#> 4 Student's t-test
#> 5 Student's t-test
#> 6 Student's t-testWithin-subjects design
# for reproducibility
set.seed(123)
# parametric
pairwise_comparisons(
data = bugs_long,
x = condition,
y = desire,
subject.id = subject,
type = "parametric",
paired = TRUE,
p.adjust.method = "BH"
)
#> # A tibble: 6 x 6
#> group1 group2 p.value test.details p.value.adjustment
#> <chr> <chr> <dbl> <chr> <chr>
#> 1 HDHF HDLF 1.06e- 3 Student's t-test FDR
#> 2 HDHF LDHF 7.02e- 2 Student's t-test FDR
#> 3 HDHF LDLF 3.95e-12 Student's t-test FDR
#> 4 HDLF LDHF 6.74e- 2 Student's t-test FDR
#> 5 HDLF LDLF 1.99e- 3 Student's t-test FDR
#> 6 LDHF LDLF 6.66e- 9 Student's t-test FDR
#> label
#> <chr>
#> 1 list(~italic(p)[FDR-corrected]==0.001)
#> 2 list(~italic(p)[FDR-corrected]==0.070)
#> 3 list(~italic(p)[FDR-corrected]==3.95e-12)
#> 4 list(~italic(p)[FDR-corrected]==0.067)
#> 5 list(~italic(p)[FDR-corrected]==0.002)
#> 6 list(~italic(p)[FDR-corrected]==6.66e-09)
# non-parametric
pairwise_comparisons(
data = bugs_long,
x = condition,
y = desire,
subject.id = subject,
type = "nonparametric",
paired = TRUE,
p.adjust.method = "BY"
)
#> # A tibble: 6 x 11
#> group1 group2 statistic p.value alternative
#> <chr> <chr> <dbl> <dbl> <chr>
#> 1 HDHF HDLF 4.78 1.44e- 5 two.sided
#> 2 HDHF LDHF 2.44 4.47e- 2 two.sided
#> 3 HDHF LDLF 8.01 5.45e-13 two.sided
#> 4 HDLF LDHF 2.34 4.96e- 2 two.sided
#> 5 HDLF LDLF 3.23 5.05e- 3 two.sided
#> 6 LDHF LDLF 5.57 4.64e- 7 two.sided
#> method
#> <chr>
#> 1 Durbin's all-pairs test for a two-way balanced incomplete block design
#> 2 Durbin's all-pairs test for a two-way balanced incomplete block design
#> 3 Durbin's all-pairs test for a two-way balanced incomplete block design
#> 4 Durbin's all-pairs test for a two-way balanced incomplete block design
#> 5 Durbin's all-pairs test for a two-way balanced incomplete block design
#> 6 Durbin's all-pairs test for a two-way balanced incomplete block design
#> distribution p.adjustment test.details p.value.adjustment
#> <chr> <chr> <chr> <chr>
#> 1 t none Durbin-Conover test BY
#> 2 t none Durbin-Conover test BY
#> 3 t none Durbin-Conover test BY
#> 4 t none Durbin-Conover test BY
#> 5 t none Durbin-Conover test BY
#> 6 t none Durbin-Conover test BY
#> label
#> <chr>
#> 1 list(~italic(p)[BY-corrected]==1.44e-05)
#> 2 list(~italic(p)[BY-corrected]==0.045)
#> 3 list(~italic(p)[BY-corrected]==5.45e-13)
#> 4 list(~italic(p)[BY-corrected]==0.050)
#> 5 list(~italic(p)[BY-corrected]==0.005)
#> 6 list(~italic(p)[BY-corrected]==4.64e-07)
# robust
pairwise_comparisons(
data = bugs_long,
x = condition,
y = desire,
subject.id = subject,
type = "robust",
paired = TRUE,
p.adjust.method = "hommel"
)
#> # A tibble: 6 x 11
#> group1 group2 estimate conf.level conf.low conf.high p.value p.crit
#> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 HDHF HDLF 1.03 0.95 0.140 1.92 0.00999 0.0127
#> 2 HDHF LDHF 0.454 0.95 -0.104 1.01 0.0520 0.025
#> 3 HDHF LDLF 1.95 0.95 1.09 2.82 0.000000564 0.00851
#> 4 HDLF LDHF -0.676 0.95 -1.61 0.256 0.0520 0.05
#> 5 HDLF LDLF 0.889 0.95 0.0244 1.75 0.0203 0.0169
#> 6 LDHF LDLF 1.35 0.95 0.560 2.14 0.000102 0.0102
#> test.details p.value.adjustment
#> <chr> <chr>
#> 1 Yuen's trimmed means test Hommel
#> 2 Yuen's trimmed means test Hommel
#> 3 Yuen's trimmed means test Hommel
#> 4 Yuen's trimmed means test Hommel
#> 5 Yuen's trimmed means test Hommel
#> 6 Yuen's trimmed means test Hommel
#> label
#> <chr>
#> 1 list(~italic(p)[Hommel-corrected]==0.010)
#> 2 list(~italic(p)[Hommel-corrected]==0.052)
#> 3 list(~italic(p)[Hommel-corrected]==5.64e-07)
#> 4 list(~italic(p)[Hommel-corrected]==0.052)
#> 5 list(~italic(p)[Hommel-corrected]==0.020)
#> 6 list(~italic(p)[Hommel-corrected]==1.02e-04)
# Bayesian
pairwise_comparisons(
data = bugs_long,
x = condition,
y = desire,
subject.id = subject,
type = "bayes",
paired = TRUE,
bf.prior = 0.77
)
#> # A tibble: 6 x 18
#> group1 group2 term estimate conf.level conf.low conf.high pd
#> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 HDHF HDLF Difference -1.10 0.95 -1.73 -0.492 1
#> 2 HDHF LDHF Difference -0.465 0.95 -0.969 0.0406 0.962
#> 3 HDHF LDLF Difference -2.13 0.95 -2.64 -1.63 1
#> 4 HDLF LDHF Difference 0.652 0.95 -0.0362 1.32 0.971
#> 5 HDLF LDLF Difference -0.983 0.95 -1.60 -0.423 0.999
#> 6 LDHF LDLF Difference -1.67 0.95 -2.14 -1.14 1
#> rope.percentage prior.distribution prior.location prior.scale bf10
#> <dbl> <chr> <dbl> <dbl> <dbl>
#> 1 0 cauchy 0 0.77 3.95e+ 1
#> 2 0.170 cauchy 0 0.77 5.42e- 1
#> 3 0 cauchy 0 0.77 1.22e+10
#> 4 0.162 cauchy 0 0.77 6.50e- 1
#> 5 0 cauchy 0 0.77 1.72e+ 1
#> 6 0 cauchy 0 0.77 4.78e+ 6
#> method log_e_bf10 expression label
#> <chr> <dbl> <list> <chr>
#> 1 Bayesian t-test 3.68 <language> list(~log[e](BF['01'])==-3.68)
#> 2 Bayesian t-test -0.612 <language> list(~log[e](BF['01'])==0.61)
#> 3 Bayesian t-test 23.2 <language> list(~log[e](BF['01'])==-23.22)
#> 4 Bayesian t-test -0.430 <language> list(~log[e](BF['01'])==0.43)
#> 5 Bayesian t-test 2.84 <language> list(~log[e](BF['01'])==-2.84)
#> 6 Bayesian t-test 15.4 <language> list(~log[e](BF['01'])==-15.38)
#> test.details
#> <chr>
#> 1 Student's t-test
#> 2 Student's t-test
#> 3 Student's t-test
#> 4 Student's t-test
#> 5 Student's t-test
#> 6 Student's t-test
Using {pairwiseComparisons} with ggsignif
Example-1: between-subjects
# needed libraries
set.seed(123)
library(ggplot2)
library(pairwiseComparisons)
library(ggsignif)
# converting to factor
mtcars$cyl <- as.factor(mtcars$cyl)
# creating a basic plot
p <- ggplot(mtcars, aes(cyl, wt)) +
geom_boxplot()
# using `{pairwiseComparisons}` package to create a dataframe with results
set.seed(123)
(df <-
pairwise_comparisons(mtcars, cyl, wt) %>%
dplyr::mutate(groups = purrr::pmap(.l = list(group1, group2), .f = c)) %>%
dplyr::arrange(group1))
#> # A tibble: 3 x 12
#> group1 group2 statistic p.value alternative method distribution
#> <chr> <chr> <dbl> <dbl> <chr> <chr> <chr>
#> 1 4 6 5.39 0.00831 two.sided Games-Howell test q
#> 2 4 8 9.11 0.0000124 two.sided Games-Howell test q
#> 3 6 8 5.12 0.00831 two.sided Games-Howell test q
#> p.adjustment test.details p.value.adjustment
#> <chr> <chr> <chr>
#> 1 none Games-Howell test Holm
#> 2 none Games-Howell test Holm
#> 3 none Games-Howell test Holm
#> label groups
#> <chr> <list>
#> 1 list(~italic(p)[Holm-corrected]==0.008) <chr [2]>
#> 2 list(~italic(p)[Holm-corrected]==1.24e-05) <chr [2]>
#> 3 list(~italic(p)[Holm-corrected]==0.008) <chr [2]>
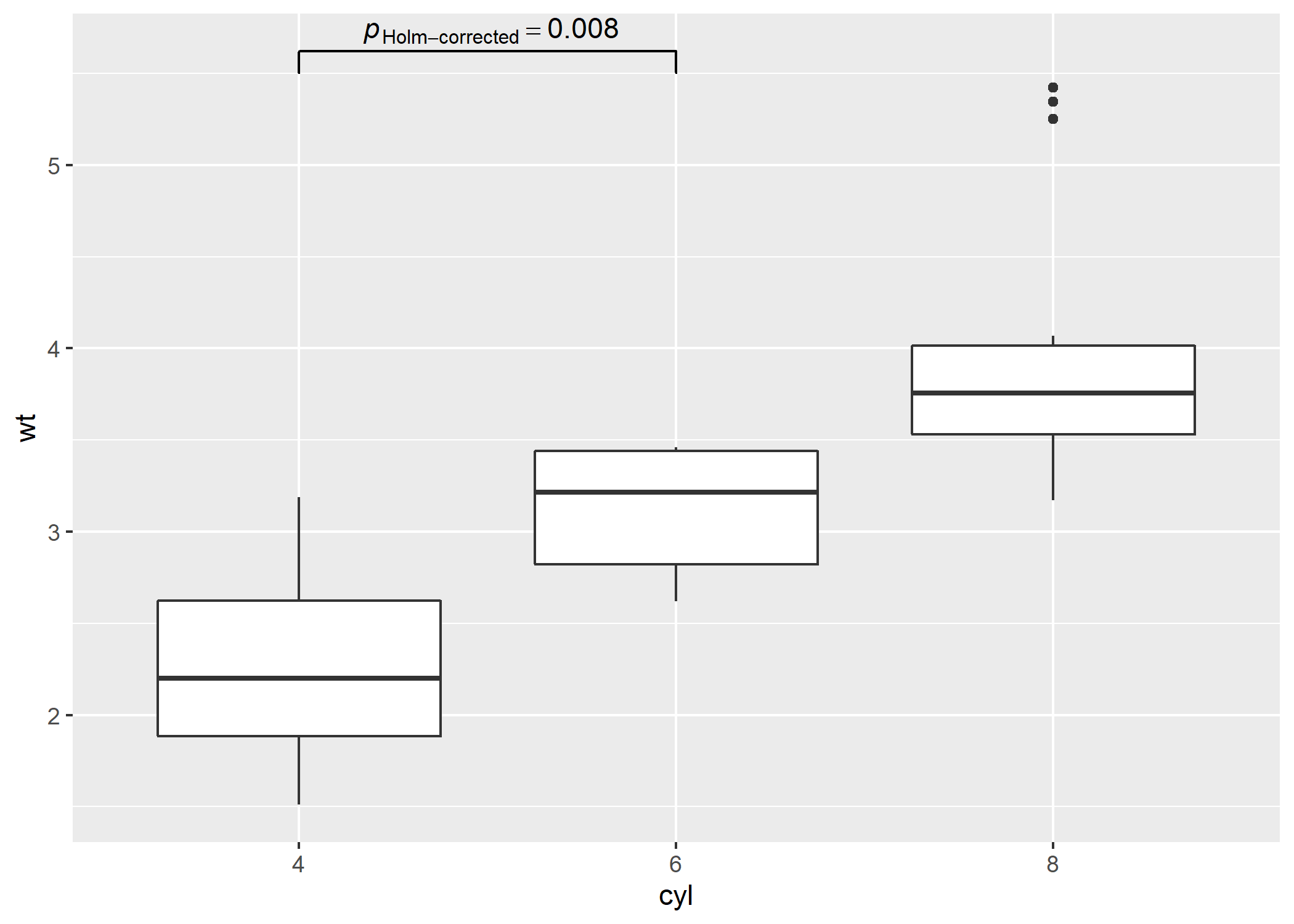
# using `geom_signif` to display results
# (note that you can choose not to display all comparisons)
p +
ggsignif::geom_signif(
comparisons = list(df$groups[[1]]),
annotations = df$label[[1]],
test = NULL,
na.rm = TRUE,
parse = TRUE
)
Example-2: within-subjects
# needed libraries
library(ggplot2)
library(pairwiseComparisons)
library(ggsignif)
# creating a basic plot
p <- ggplot(WRS2::WineTasting, aes(Wine, Taste)) +
geom_boxplot()
# using `{pairwiseComparisons}` package to create a dataframe with results
set.seed(123)
(df <-
pairwise_comparisons(
WRS2::WineTasting,
Wine,
Taste,
subject.id = Taster,
type = "bayes",
paired = TRUE
) %>%
dplyr::mutate(groups = purrr::pmap(.l = list(group1, group2), .f = c)) %>%
dplyr::arrange(group1))
#> # A tibble: 3 x 19
#> group1 group2 term estimate conf.level conf.low conf.high pd
#> <chr> <chr> <chr> <dbl> <dbl> <dbl> <dbl> <dbl>
#> 1 Wine A Wine B Difference -0.00721 0.95 -0.0569 0.0404 0.624
#> 2 Wine A Wine C Difference -0.0766 0.95 -0.140 -0.0144 0.989
#> 3 Wine B Wine C Difference -0.0696 0.95 -0.109 -0.0330 1.00
#> rope.percentage prior.distribution prior.location prior.scale bf10
#> <dbl> <chr> <dbl> <dbl> <dbl>
#> 1 0.404 cauchy 0 0.707 0.235
#> 2 0.000526 cauchy 0 0.707 3.71
#> 3 0 cauchy 0 0.707 50.5
#> method log_e_bf10 expression label
#> <chr> <dbl> <list> <chr>
#> 1 Bayesian t-test -1.45 <language> list(~log[e](BF['01'])==1.45)
#> 2 Bayesian t-test 1.31 <language> list(~log[e](BF['01'])==-1.31)
#> 3 Bayesian t-test 3.92 <language> list(~log[e](BF['01'])==-3.92)
#> test.details groups
#> <chr> <list>
#> 1 Student's t-test <chr [2]>
#> 2 Student's t-test <chr [2]>
#> 3 Student's t-test <chr [2]>
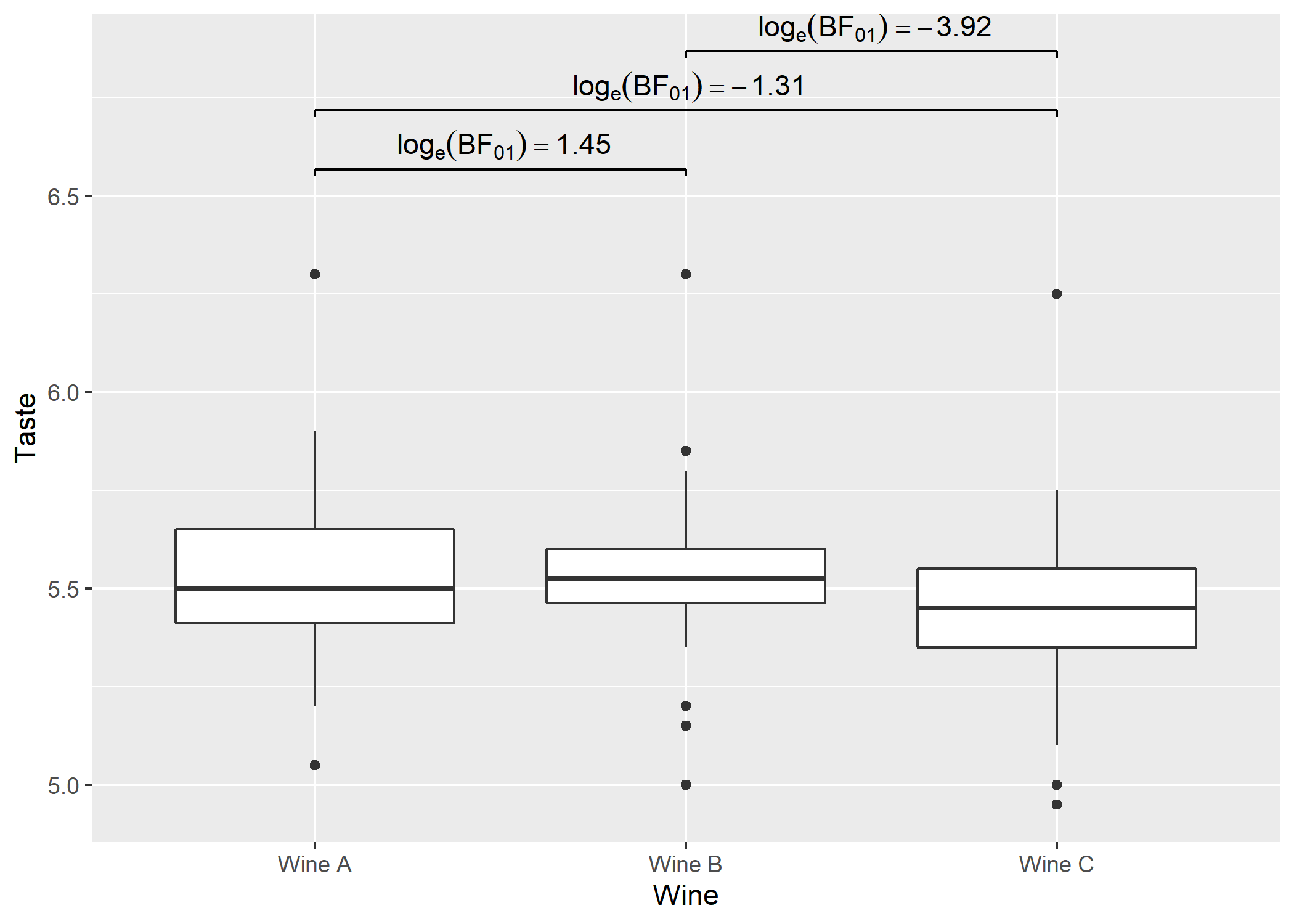
# using `geom_signif` to display results
p +
ggsignif::geom_signif(
comparisons = df$groups,
map_signif_level = TRUE,
tip_length = 0.01,
y_position = c(6.5, 6.65, 6.8),
annotations = df$label,
test = NULL,
na.rm = TRUE,
parse = TRUE
)